- B
- K
- P
Hello! We are 3 biomolecular engineering students at University of California, Santa Cruz (UCSC) completing our senior academic year-long capstone series (BME 129). We would love support from the community to raise funds to develop a novel genetic engineering tool that will benefit a wide range of scientific and medical fields.
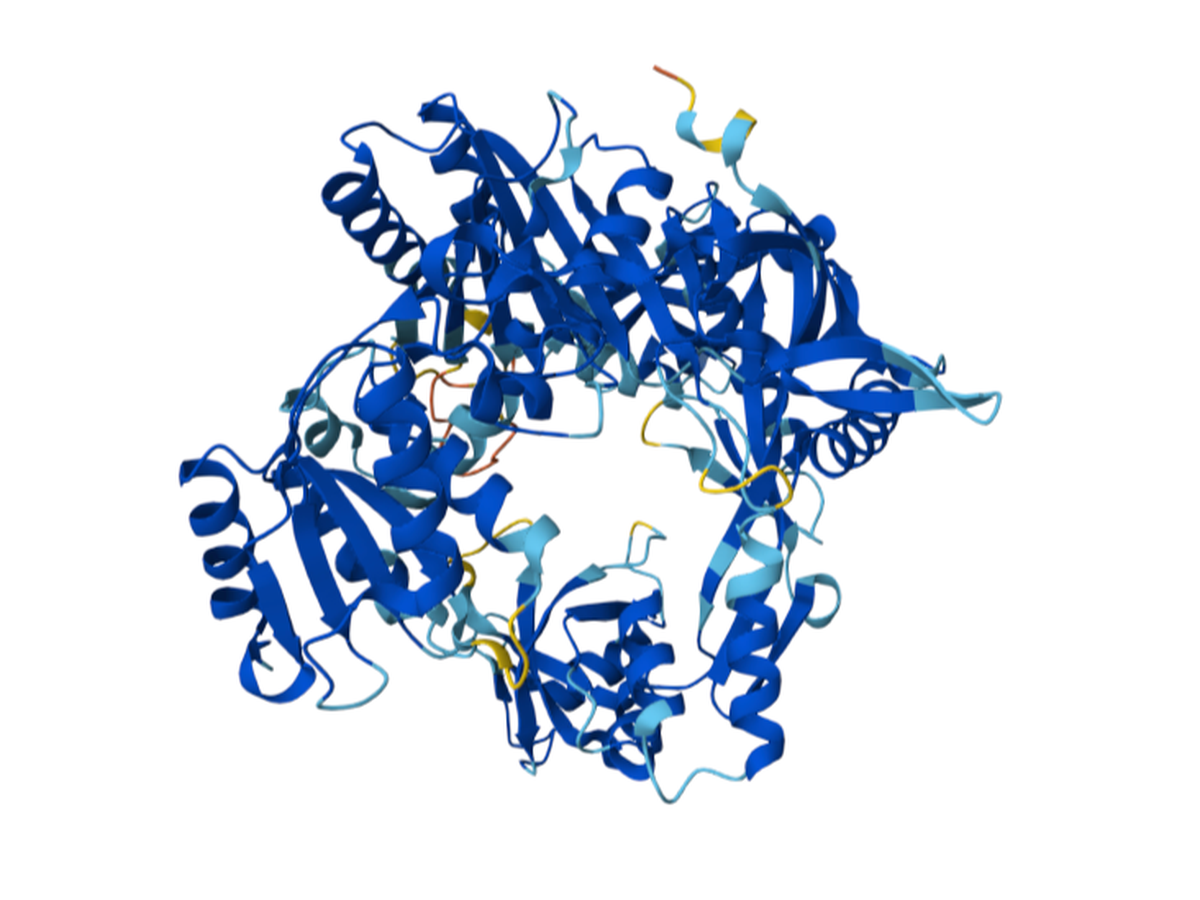
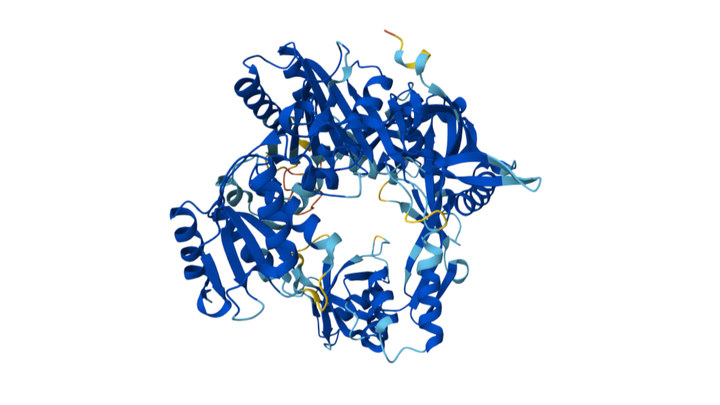
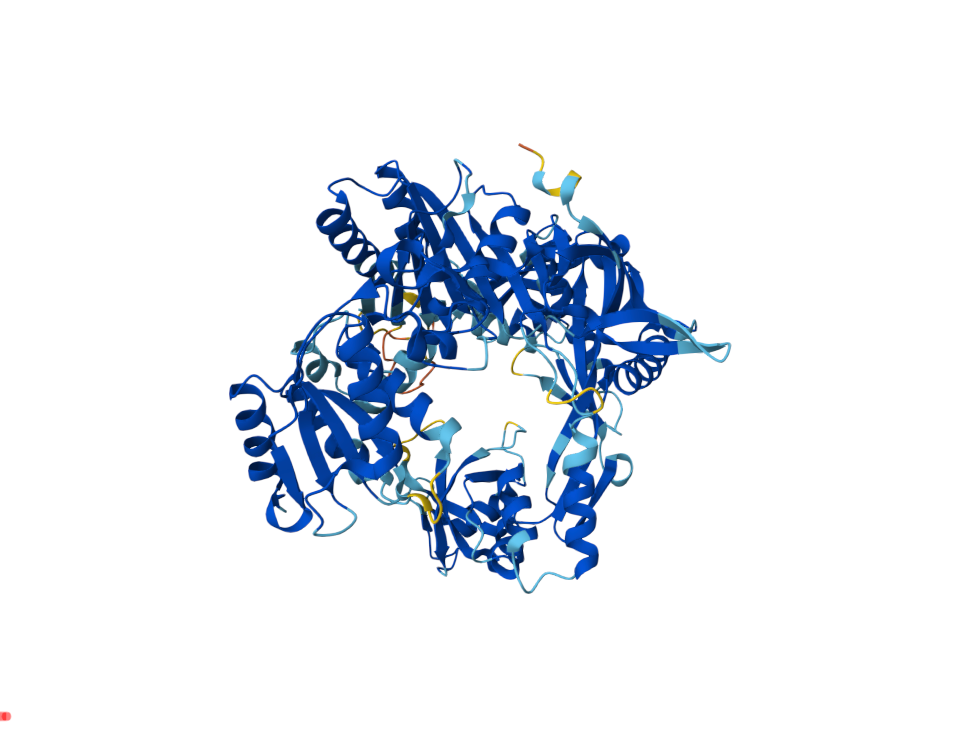
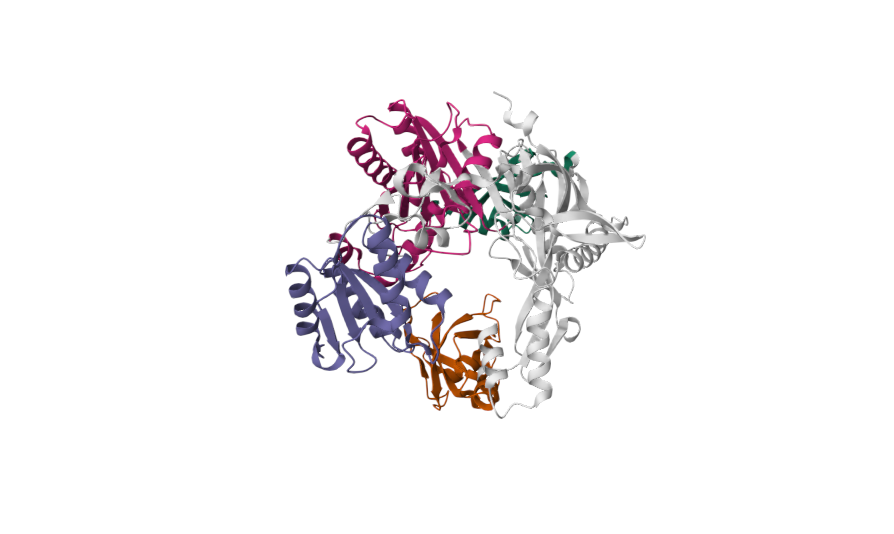
Current bacterial gene-editing technologies often rely on complex RNA-based systems like CRISPR-Cas, which can be difficult to adapt across all prokaryotes. Prokaryotic Argonaute proteins (pAgos) can serve as part of an organism's immune system. Naturally, they exist within cells to protect against invading genetic material by using guide DNA or RNA to cleave complementary sequences on the invading genetic element.
Synechococcus elongatus, a highly researched cyanobacterium, serves as an appealing organism to study because of its mesophilic nature (~20-45°C). This makes the Argonaute protein of S. elongatus fundamentally compatible with common laboratory organisms such as E. coli. Normally, Synechococcus elongatus (SeAgo) performs single-strand breaks (SSBs), but codon optimization can be utilized to create a programmable DNA-cleaving tool capable of generating targeted double-strand breaks (DSBs) on genomic DNA. This method of genetic modification provides a more flexible alternative to CRISPR-Cas protein systems since they're limited to cleaving specifically near sequence motifs (PAM sequences) and depend on RNA intermediates.
When expressed in E. coli, SeAgo can be directed with synthetic DNA guides to generate precise DNA fragments, allowing for targeted genome modification without reliance on RNA intermediates. This DNA-guided mechanism provides greater flexibility and sustainability measures because of how cost-effective guide DNA is to synthesize. By harnessing SeAgo’s unique properties, this project aims to expand the molecular biology repertoire for microbial genetic engineering and synthetic biology applications.